Copper-Free Click Chemistry Visualizes Plant Cell Wall Dynamics
November 18, 2025
In a pioneering study titled "Dynamic imaging of cell wall polysaccharides by metabolic click-mediated labeling of pectins in living elongating cells" published in The Plant Journal (2022), Ropitaux and colleagues address the long-standing challenge of visualizing real-time dynamics of plant cell wall polysaccharides, which are critical for growth and environmental interactions but inaccessible to traditional fixation-dependent methods like immunolabeling. The research introduces an innovative two-step methodology combining metabolic incorporation of azide-functionalized sugar analogs (e.g., Kdo-N3 for rhamnogalacturonan-II and Fuc-N3 for pectins) with copper-free strain-promoted azide-alkyne cycloaddition (SPAAC) to enable non-invasive, fluorescent tagging of glycans in actively elongating cells. This approach successfully captures dynamic deposition patterns—such as "stop-and-go" growth segments in tobacco pollen tubes—and reveals spatial organization of pectins in Arabidopsis roots, all while maintaining full cell viability. By transitioning from static snapshots to live, real-time imaging, the study unlocks new avenues for investigating polysaccharide remodeling, gene function, and environmental responses in plant biology.
The Core Challenge: Visualizing Living Glycans
Unlike proteins, which can be tagged with fluorescent proteins like GFP and tracked in real-time, complex glycans are not directly encoded by genes. They are synthesized by the coordinated action of enzymes within the cell's biosynthetic machinery. This fundamental difference meant that observing the deposition and remodeling of polysaccharides like pectin in a living, growing cell was previously out of reach. The study by Ropitaux et al. addresses this gap head-on by leveraging the power of bio-orthogonal "click chemistry."
The Revolutionary Approach: Metabolic Labeling and Copper-Free Click Chemistry
The methodology is elegantly based on a two-step process: metabolic incorporation and fluorescent tagging.
Growing cells, such as pollen tubes and Arabidopsis roots, are fed with bio-orthogonal sugar analogs—monosaccharides that are chemically "clickable" but still recognizable by the cell's own enzymes. The researchers used two key analogs:
- Kdo-N3 (8-azido-3-deoxy-d-manno-oct-2-ulosonic acid): An azide-modified version of Kdo, a sugar specifically incorporated into the sophisticated pectic polysaccharide Rhamnogalacturonan-II (RG-II).
- Fuc-N3 (6-azido-6-deoxy-l-galactose): An azide-modified fucose, which is incorporated into pectins (like Rhamnogalacturonan-I, RG-I) and other glycans.
As the cell grows, it unknowingly uses these "tagged" sugars to build new cell wall polymers, seamlessly incorporating the azide chemical reporters into the architecture of the wall.
- Copper-Free Click Labeling (SPAAC)
This is the critical innovation that preserves cell viability. Instead of the classic copper-catalyzed click reaction, which is toxic to cells, the team employed strain-promoted azide-alkyne cycloaddition (SPAAC). This "copper-free" click reaction involves a strained alkyne probe (e.g., sDIBO) conjugated to a fluorophore. The strained alkyne reacts rapidly and specifically with the azide groups previously incorporated into the cell wall, attaching the fluorescent tag directly to the polysaccharides.
The major advantage of SPAAC is the absence of toxic copper catalysts. This allows the chemical ligation to occur under physiological conditions, keeping the cells fully alive and capable of continued growth throughout the labeling and subsequent imaging process.
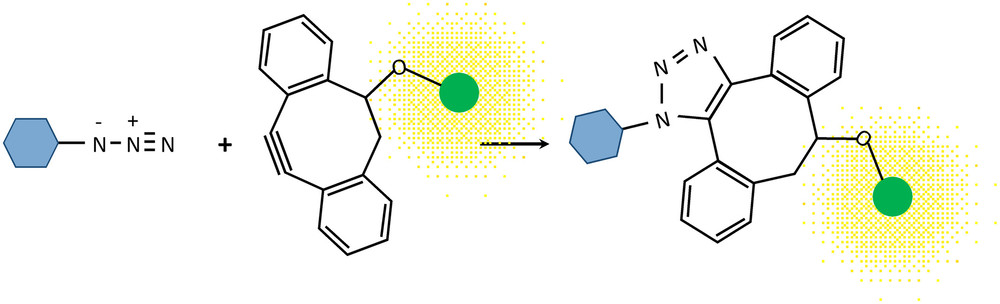 Fig.1 Mechanism of copper-free click labeling. (Ropitaux, et al., 2022)
Fig.1 Mechanism of copper-free click labeling. (Ropitaux, et al., 2022)
- A Proof of Concept in Fast-Growing Cells
The researchers meticulously developed and validated their protocol using tobacco pollen tubes, which are ideal models due to their extremely fast, tip-polarized growth.
The process involves germinating pollen grains, incubating them with the azido sugars (Kdo-N3 or Fuc-N3) to allow for metabolic incorporation, washing away the excess, and then performing the SPAAC reaction with the fluorescent probe.
The power of this technique was demonstrated by observing different labeling patterns directly correlated with the growth history of individual pollen tubes. For instance, tubes that grew only during the incorporation phase were labeled along their entire length. In contrast, tubes that continued to grow after the incorporation phase showed a distinct "stop-and-go" pattern: a brightly labeled segment corresponding to growth during the feeding period, followed by a non-fluorescent segment representing new growth after the azido sugar was removed. This visually captures the dynamic deposition of cell wall material in real-time.
Crucially, the study confirmed that neither the azido sugars nor the SPAAC reaction affected pollen tube viability or growth rates, underscoring the non-invasive nature of the technique.
Expanding the Scope to Roots and Future Applications
The methodology was successfully extended to Arabidopsis thaliana roots, another critical model system. Here, SPAAC labeling with Kdo-N3 revealed the abundant presence of RG-II at the apex of emerging and growing root hairs, as well as in the anticlinal walls of root cells in the differentiation zone. This spatial information provides new insights into the role of specific pectins in root development.
The authors highlight future directions that are now possible with this technique:
- Long-term dynamics: Studying polysaccharide remodeling and potential recycling over time.
- Super-resolution imaging: Coupling SPAAC with techniques like STED microscopy to achieve nanoscale resolution of polysaccharide organization.
- Functional studies: Applying this method to Arabidopsis mutants defective in cell wall biosynthesis or remodeling to understand gene function.
- Environmental responses: Using pulse-labeling to investigate how stresses (e.g., nutrient deficiency, heat) specifically alter cell wall biosynthesis and modification during the treatment period.
Bridging Pioneering Research with Advanced Glycobiology Solutions
The work by Ropitaux et al. is a testament to the power of click chemistry in plant glycobiology. It moves the field from static observations to dynamic, functional studies of the cell wall in its native, living state.
At CD BioGlyco, our GlycoCLICK™ platform is built upon these very principles. We offer a comprehensive suite of services that mirror and extend the capabilities demonstrated in this seminal paper:
- GlycoCLICK™-based Imaging Service: Our specialists can assist in designing and implementing labeling strategies for your specific cell types or tissues, leveraging techniques like SPAAC for live-cell imaging.
- GlycoCLICK™-based Drug and Vaccine Development: The ability to label and target specific glycans opens doors for developing glycan-based inhibitors, drug delivery systems, and conjugate vaccines.
The methodology outlined by Ropitaux et al. is no longer confined to pioneering labs. Through our GlycoCLICK™ technology and expert team, CD BioGlyco empowers researchers worldwide to integrate these advanced techniques into their own projects, accelerating discovery in plant science, biochemistry, and biomedical research. Unlock the dynamics of your glycobiology research. Contact us today to learn how our GlycoCLICK™ services can bring your projects to life.
Reference
- Ropitaux, M.; et al. Dynamic imaging of cell wall polysaccharides by metabolic click-mediated labeling of pectins in living elongating cells. The Plant Journal. 2022, 110(3): 916-924.

 Fig.1 Mechanism of copper-free click labeling. (Ropitaux, et al., 2022)
Fig.1 Mechanism of copper-free click labeling. (Ropitaux, et al., 2022)